Second Edition (2025) will be out in October 2025
Check my blog on the Cambridge website
Fifty Years of International Environmental Law: Looking Back and Looking Ahead

Second Edition (2025) will be out in October 2025
Check my blog on the Cambridge website
Fifty Years of International Environmental Law: Looking Back and Looking Ahead
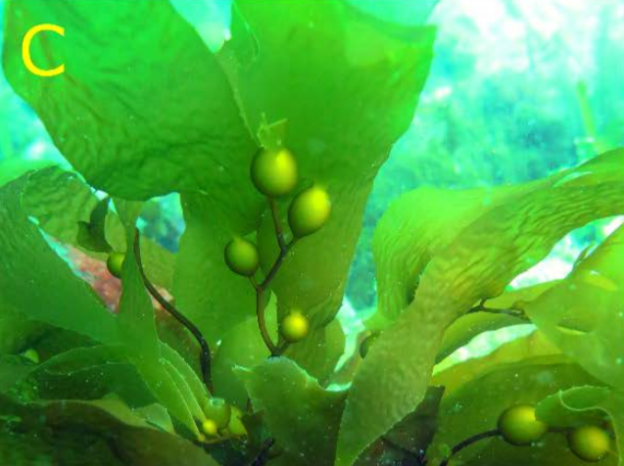

At a salmon farm in Tasmania Australia is an experiment that researchers hope can save an entire ecosystem from warming oceans. Beneath the waves, scientists are growing several types of giant kelp—which in the wild can grow up to 175 feet tall—on rope to track which ones can thrive in hotter conditions. Rising water temperatures, more frequent marine heat waves and invasive sea urchins have already destroyed some 95% of the giant kelp forests in Tasmania, scientists say. The island south of Australia’s mainland is a global hot spot for ocean warming, with sea temperatures in the island’s east rising faster than the global average, a dynamic that has already wreaked havoc on some marine species in a place where fishing remains a key industry.
In Tasmania, scientists are conducting experiments to identify heat-tolerant giant kelp, plan to use artificial intelligence and genetic analysis to better understand why some types fare better than others, and will then have to figure out a way to plant them in the wild without it costing a fortune. Eventually, they could use the genetic information to breed kelp to be even more heat tolerant…But success is far from guaranteed. Running lab tests on kelp can be tricky, given the great size the plants can reach. Efforts to control the invasive long-spined sea urchins, including with government subsidies that encourage fishermen to catch them, could fail. Marine heat waves could increase beyond the ability of any kelp to cope. Scientists also still aren’t sure to what extent genetic factors allow giant kelp to survive in warmer water, or whether environmental factors—such as nutrient and light availability—are more important.
Excerpts from Mike Cherney, Inside the Quest for a Super Kelp That Can Survive Hotter, WSJ, Feb. 22, 2024
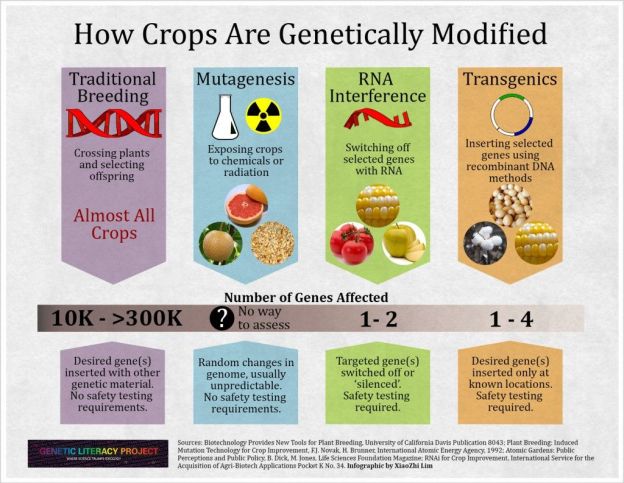
China is betting that CRISP technology*can transform the country’s food supply. China also expanded its efforts beyond its borders in 2017, when the state-owned company ChemChina bought Switzerland-based Syngenta—one of the world’s four largest agribusinesses, which has a large R&D team working with CRISPR—for $43 billion. That was the most China has ever spent on acquiring a foreign company, and it created an intimate relationship between government, industry, and academia—a “sort of a ménage à trois” that ultimately could funnel intellectual property from university labs into the company, says plant geneticist Zachary Lippman of Cold Spring Harbor Laboratory in New York.
Chinese leaders “want to strategically invest in genome editing, and [by that] I mean, catch up,” says Zhang Bei, who heads a team of 50 scientists at the Syngenta Beijing Innovation Center…China may one day need CRISPR-modified plants to provide enough food for its massive population…. China needs to resolve how it will regulate CRISPR-engineered crops—a divisive issue in many countries. In a 2018 decision that rocked big agriculture, a European court ruled that such crops are genetically modified organisms (GMOs) that need strict regulation. In contrast, the U.S. Department of Agriculture (USDA) exempts genome-edited plants from regulations covering GMOs as long as they were produced not by transferring DNA from other species, but by inducing mutations that could have occurred naturally or through conventional breeding. Chinese consumers are wary of GM food. The country strictly limits the import of GM crops, and the only GM food it grows are papayas for domestic consumption. But for CRISPR, many plant researchers around assume China will follow in the United States’s footsteps…
For Corteva, Syngenta, and the other two big ag companies—BASF and Bayer (which acquired Monsanto last year)—the long game is to use CRISPR to develop better versions of their serious moneymakers, the “elite” varieties of a wide range of crops that have big commercial markets. They sell dozens of kinds of elite corn seeds—for example, inbred strains that consistently have high yields or reliable resistance to herbicides. Creating the genetic purity needed for an elite variety typically takes traditional breeding of many generations of plants, and CRISPR is seen as the cleanest way to improve them quickly. The earlier methods of engineering a plant can lead to unwanted genomic changes that must be laboriously culled…
Syngenta sees CRISPR-modified corn as a big opportunity in China, which grows more hectares of corn than any other crop. Yields per hectare are only 60% of those in the United States because corn ear worms often weaken Chinese crops. A fungus thrives in the weakened plants, producing a toxin that makes the resultant ears unfit for animal feed. As a result, China must import a great deal of corn. (According to USDA, 82% of U.S.-grown corn has been engineered to have a bacterial gene that makes it resistant to ear worms.)…“Syngenta is putting a lot of emphasis to grow in China to become the leading seed company. The China market as a whole, if it modernizes as the U.S. has modernized, can be as big as the U.S. market.”
Jon Cohen, To feed its 1.4 billion, China bets big on genome editing of crops, Science Magazine, Aug. 2, 2019
* Genome editing (also called gene editing) is a group of technologies that give scientists the ability to change an organism’s DNA. These technologies allow genetic material to be added, removed, or altered at particular locations in the genome. Several approaches to genome editing have been developed. A recent one is known as CRISPR-Cas9.

Breakthroughs in the science of programmable gene expression inspired DARPA to establish the PReemptive Expression of Protective Alleles and Response Elements (PREPARE) program with the goal of delivering powerful new defenses against public health and national security threats. DARPA has now selected five teams to develop a range of new medical interventions that temporarily and reversibly modulate the expression of protective genes to guard against acute threats from influenza and ionizing radiation, which could be encountered naturally, occupationally, or through a national security event.
The program builds from the understanding that the human body has innate defenses against many types of health threats, but that the body does not always activate these defenses quickly or robustly enough to block the worst damage. To augment existing physiological responses, PREPARE technologies would provide a programmable capability to up- or down-regulate gene expression on demand, providing timely, scalable defenses that are proportional to anticipated threats. Service members and first responders could administer these interventions prior to threat exposure or therapeutically after exposure to mitigate the risk of harm or death.

Influenza: “Researchers working within the PREPARE program seek to improve rates of survival and recovery in catastrophic scenarios for which reliable and scalable countermeasures don’t currently exist,” said Dr. Renee Wegrzyn, the PREPARE program manager….Three PREPARE teams are pursuing multi-pronged approaches to influenza defense and treatment that use programmable gene modulators to boost the human body’s natural defenses against influenza and also weaken the virus’ ability to cause harm by directly neutralizing the viral genomes. If successful, their approaches would potentially protect against virtually all influenza strains — regardless of whether a virus is newly emergent or has developed drug resistance — and would provide near instantaneous immunity, in contrast to traditional vaccines. Additionally, the teams are designing their countermeasures so that they are simple to deliver — for example, as intranasal sprays — reducing the logistical challenge of protecting large numbers of people.A team led by DNARx LLC, under principal investigator Dr. Robert Debs, aims to develop a new DNA-encoded gene therapy that helps patients fight influenza by boosting the natural immune response and other protective functions of their nasal passages and lungs.

Ionizing Gamma Radiation: Other PREPARE teams are pursuing treatments to protect the body from the effects of ionizing gamma radiation. In humans, radiation poisoning primarily affects stem cells in the blood and gut, yet existing treatments only help to regenerate blood cells, and only with limited effect. There is no possibility for prophylactic administration of these drugs, and most must be delivered immediately following radiation exposure to provide any benefit. There are no existing medical countermeasures for radiation damage to the gut…
A team led by the University of California, San Francisco, under principal investigator Dr. Jonathan Weissman, also aims to develop gene therapies to enhance resilience against ionizing radiation. The team’s approach should result in an intravenous or orally available treatment that activates innate defenses in gut and blood stem cells for a period of several weeks.
A Dose of Inner Strength to Survive and Recover from Potentially Lethal Health Threats
New tools for programmable modulation of gene expression could yield enhanced resilience against influenza and ionizing radiation for service members and first responders, DARPA Press Release, June 27, 2019
In the first attempt of its kind, researchers plan to sequence all known species of eukaryotic life—66,000 species of animals, plants, fungi, and protozoa—in a single country, the United Kingdom. The announcement was made here today at the official launch of an even grander $4.7 billion global effort, called the Earth BioGenome Project (EBP), to sequence the genomes of all of Earth’s known 1.5 million species of eukaryotes within a decade.
In terms of genomes sequenced, the eukaryotes—the branch of complex life consisting of organisms with cells that have a nucleus inside a membrane—lag far behind the bacteria and archaea. Researchers have sequenced just about 3500 eukaryotic genomes, and only 100 at high quality.
The U.K. sequencing effort—dubbed The Darwin Tree of Life project—will now become part of the EBP mix…Also announced today was a memorandum of understanding for participating in EBP. It has been signed by 19 institutions, including BGI Shenzhen, China; the Royal Botanic Gardens, Kew; and Sanger.
Excerpts from Erik Stokstad, Researchers launch plan to sequence 66,000 species in the United Kingdom. But that’s just a start, Science, Nov. 1, 2018
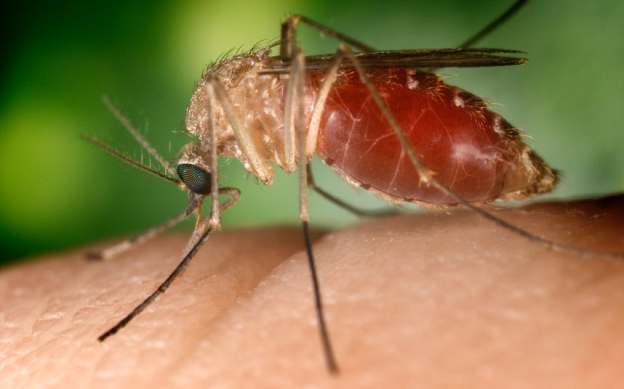
The mosquitoes are being fitted with a piece of dna called a gene drive. Unlike the genes introduced into run-of-the-mill genetically modified organisms, gene drives do not just sit still once inserted into a chromosome. They actively spread themselves, thereby reaching more and more of the population with each generation. If their effect is damaging, they could in principle wipe out whole species.. If gene drives were to condemn to a similar fate the mosquitoes that spread malaria, a second of humankind’s great scourges might be consigned to history.
Gene drives can in principle be used against any creatures which reproduce sexually with short generations and aren’t too rooted to a single spot. The insects that spread leishmaniasis, Chagas disease, dengue fever, chikungunya, trypanosomiasis and Zika could all be potential targets. So could creatures which harm only humankind’s dominion, not people themselves. Biologists at the University of California, San Diego, have developed a gene-drive system for Drosophila suzukii, an Asian fruitfly which, as an invasive species, damages berry and fruit crops in America and Europe. Island Conservation, an international environmental ngo, thinks gene drives could offer a humane and effective way of reversing the damage done by invasive species such as rats and stoats to native ecosystems in New Zealand and Hawaii.
Such critics fear that the laudable aim of vastly reducing deaths from malaria—which the World Health Organisation puts at 445,000 a year, most of them children—will open the door to the use of gene drives for far less clear-cut benefits in ways that will entrench some interests, such as those of industrial farmers, at the expense of others. They also point to possible military applications: gene drives could in principle make creatures that used not to spread disease more dangerous… The ability to remove species by fiat—in effect, to get them to remove themselves—is, like the prospect of making new species from scratch, a power that goes beyond the past ambit of humankind.
Gene drives based on crispr-Cas9 could easily be engineered to target specific bits of the chromosome and insert themselves seamlessly into the gap, thus ensuring that every gamete gets a copy . By 2016, gene drives had been created in yeast, fruitflies and two species of mosquito. In work published in the journal Nature Biotechnology in September, Andrea Crisanti, Mr Burt and colleagues at Imperial showed that one of their gene drives could drive a small, caged population of the mosquito Anopheles gambiae to extinction—the first time a gene drive had shown itself capable of doing this. The next step is to try this in a larger caged population.
There are also worries about how gene drives might be used to create a weapon. …The need to find ways to guard against such attacks is one of the reasons that the Pentagon’s Defence Advanced Research Projects Agency (darpa) gives for its work on gene drives. Renee Wegrzyn, programme manager for darpa’s “Safe Genes” project, says the work is to prevent “technological surprise”, whether in the form of an unintended consequence or nefarious use. One of the academic teams she funds has made progress in developing anti-crispr enzyme systems that one day might be able to inhibit a drive’s operation.
Oversight needs not just to bring together a range of government agencies; it requires co-operation between governments, too. The Cartagena Protocol on Biosafety, which entered into force under the un Convention on Biological Diversity (cbd) in 2003, provides controls on the transfer of genetically modified organisms. But how it applies to gene drives is unclear—and besides, America has never ratified the convention. An attempt to ban gene-drive research through the cbd, which was backed by the etc Group and other ngos, failed at the convention’s biennial meeting in Cancún in 2016…Like the reintroduction of vanished species advocated by the rewilding movement, gene-drive technology will provide new arenas for the fight between those who wish to defend nature and those who wish to tame it.
Excerpts from Gene Drives: Extinction on Demand, Economist, Nov. 10, 2018, at 24

Many envisioned environmental applications of newly developed gene-editing techniques such as CRISPR might provide profound benefits for ecosystems and society. But depending on the type and scale of the edit, gene-edited organisms intentionally released into the environment could also deliver off-target mutations, evolutionary resistance, ecological disturbance, and extinctions. Hence, there are ongoing conversations about the responsible application of CRISPR, especially relative to the limitations of current global governance structures to safeguard its use, Largely missing from these conversations is attention to local communities in decision-making. Most policy discussions are instead occurring at the national or international level even though local communities will be the first to feel the context-dependent impacts of any release. ..
CRISPR gene editing and other related genetic technologies are groundbreaking in their ability to precisely and inexpensively alter the genome of any species. CRISPR-based gene drives hold particular import because they are designed to rapidly spread genetic changes—including detrimental traits such as infertility—through populations of sexually reproducing organisms, to potentially reach every member of a species. Villages in Burkina Faso are weighing the release of gene drive–bearing mosquitoes that could suppress malaria. Nantucket Island residents in the United States are considering the release of genetically engineered white-footed mice to deplete Lyme disease reservoirs. New Zealand communities are discussing the possibility of using genetic methods to eliminate exotic predators.
But what if a gene drive designed to suppress an invasive species escaped its release site and spread to a native population? Or if a coral species gene edited to better adapt to environmental stressors dominated reef ecosystems at the expense of a diversity of naturally evolving coral species and the fish that depend on them ? The gravity of these potential outcomes begs the question: Should humans even be meddling with the DNA of wild organisms? The absence of generally agreed on answers can be used to support calls for moratoria on developing and releasing genetically altered organisms, especially those with gene drives (6).
However, the promising benefits of environmental gene editing cannot be dismissed. Gene drives may provide a long-sought-after tool to control vectors of infectious disease and save millions of human lives. Projects to conserve ecosystems or promote species resilience are often intended to repair human-inflicted environmental damage. Put simply, either using this technology irresponsibly or not using it at all could prove damaging to humans, our welfare, and our planet.
At the international level, the Convention on Biological Diversity (CBD) has enlisted an expert technical panel to, in part, update its Cartagena Protocol (of which the United States is not a party) that oversees transboundary transport of living modified organisms to accommodate gene drive–bearing organisms. The International Union for the Conservation of Nature (IUCN) is also developing policy to address the release of gene-edited organisms. Although the CBD and the IUCN offer fora to engage diverse public feedback, a role largely fulfilled by civil society groups, none of these agencies currently use the broad and open deliberative process we advocate….
Different societal views about the human relationship to nature will therefore shape decision-making. Local community knowledge and perspectives must therefore be engaged to address these context-dependent, value-based considerations. A special emphasis on local communities is also a matter of justice because the first and most closely affected individuals deserve a strong voice in the decision-making process…Compounding this challenge is that these decisions cannot be made in isolation. Organisms released into local environments may cross regional and even international borders. Hence, respect for and consideration of local knowledge and value systems are necessary, but insufficient, to anticipate the potentially ramifying global implications of environmental release of gene-edited organisms. What is needed is an approach that places great weight on local perspectives within a larger global vision…
The needs of ecosystems could also be given voice to inform deliberative outcomes through custodial human proxies. Inspired by legislative precedent set by New Zealand, in which the Whanganui River was granted legal “personhood,” human representatives, nominated by both an international body like the IUCN and the local community, would be responsible for upholding the health and interests of the ecosystems in question. Proposed gene-editing strategies would be placed in the larger context of alternative approaches to address the public health or environmental issue in question…
An online registry for all projects intending to release genetically engineered organisms into the environment must be created. Currently, no central database exists for environmental gene-editing applications or for decision-making outcomes associated with their deployment, and this potentially puts the global community at risk…A global coordination task force would be charged with coordinating multiple communities, nations, and regions to ensure successful deliberative outcomes. As a hypothetical example, genetic strategies to eliminate invasive possums from New Zealand must include representatives from Australia, the country likely to be affected should animals be transported outside the intended range. Similarly, the African Union is currently deliberating appropriate governance of gene drive–bearing mosquitoes to combat malaria on a regional scale.
Excerpts from Natalie Kofl et al., Editing nature: Local roots of global governance, Science Magazine, Nov. 2, 2018




The Crispr-Cas9 system consists of two main parts: an RNA guide, which scientists program to target specific locations on a genome, and the Cas9 protein, which acts as molecular scissors. The cuts trigger repairs, allowing scientists to edit DNA in the process. Think of Crispr as a cut-and-paste tool that can add or delete genetic information. Crispr can also edit the DNA of sperm, eggs and embryos—implementing changes that will be passed down to future generations. Proponents say it offers unprecedented power to direct the evolution of species.
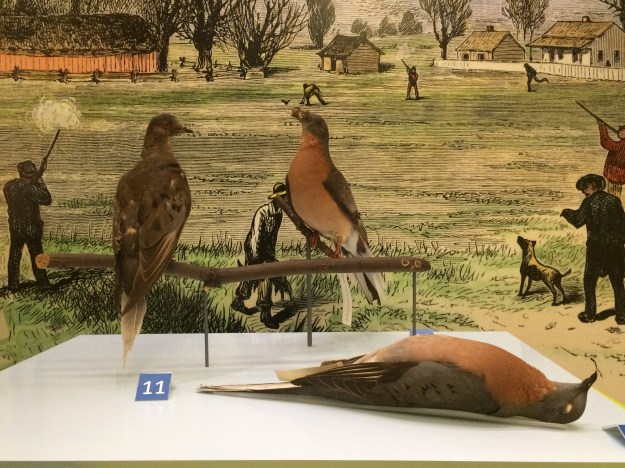
The technology is widely used in animals. Crispr has produced disease-resistant chickens and hornless dairy cattle. Scientists around the world routinely edit the genes in mice for research, adding mutations for human diseases such as autism and Alzheimer’s in a search of possible cures. Crispr-edited pigs contain kidneys that scientists hope to test as transplants in humans. Crispr has been discussed as a de-extinction tool since its earliest days. In March 2013 the conservation group Revive & Restore co-organized the first TedXDeExtinction conference in Washington, D.C. Revive & Restore was co-founded by Stewart Brand, the creator of the counterculture Whole Earth Catalog and a vocal advocate for a passenger pigeon revival.
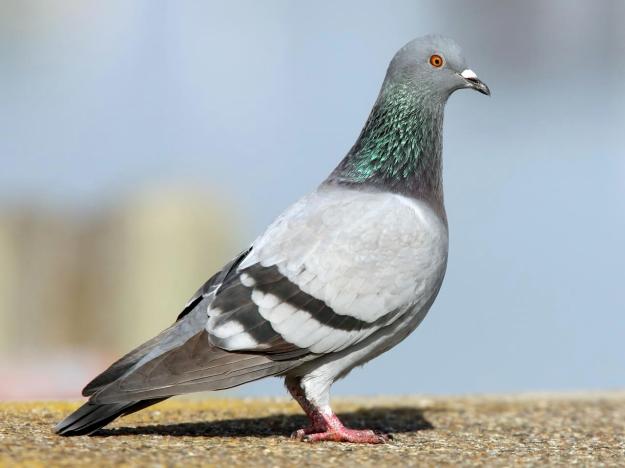
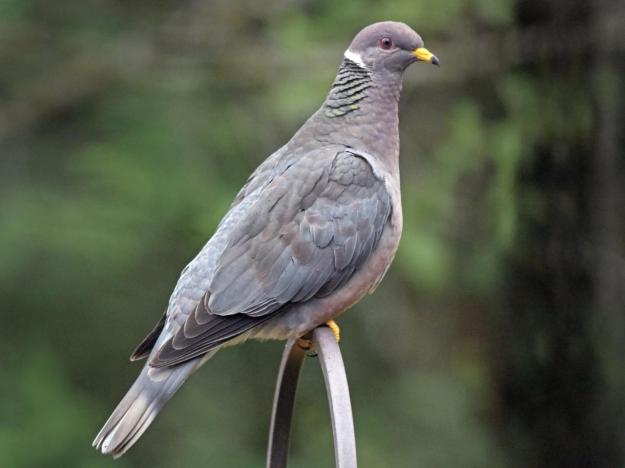
The last known passenger pigeon—a bird named Martha—died in captivity at a Cincinnati zoo in 1914….The first step was to sequence the passenger pigeon genome…Sequencing an extinct species’ genome is no easy task. When an organism dies, the DNA in its cells begins to degrade, leaving scientists with what Shapiro describes as “a soup of trillions of tiny fragments” that require reassembly. For the passenger pigeon project, Shapiro and her team took tissue samples from the toe pads of stuffed birds in museum collections. DNA in the dead tissue left them with tantalizing clues but an incomplete picture. To fill in the gaps, they sequenced the genome of the band-tailed pigeon, the passenger pigeon’s closest living relative.
By comparing the genomes of the two birds, researchers began to understand which traits distinguished the passenger pigeon. In a paper published last year in “Science,” they reported finding 32 genes that made the species unique. Some of these allowed the birds to withstand stress and disease, essential traits for a species that lived in large flocks. They found no genes that might have led to extinction. “Passenger pigeons went extinct because people hunted them to death,” Shapiro says
.Revived passenger pigeons could also face re-extinction. The species thrived in the years before European settlement of North America, when vast forests supported billions of birds. Those forests have since been replaced by cities and farmland. “The habitat the passenger pigeons need to survive is also extinct,” Shapiro says. But what does it mean to bring an extinct species back? Andre E.R. Soares, a scientist who helped sequence the passenger pigeon genome, says most people will accept a lookalike as proof of de-extinction. “If it looks like a passenger pigeon and flies like a passenger pigeon, if it has the same shape and color, they will consider it a passenger pigeon,” Soares says.
Shapiro says that’s not enough. Eventually, she says, gene-editing tools may be able to create a genetic copy of an extinct species, “but that doesn’t mean you are going to end up with an animal that behaves like a passenger pigeon or a woolly mammoth.” We can understand the nature of an extinct species through its genome, but nurture is another matter.
After he determines how passenger pigeon DNA manifests in the rock pigeons, Novak hopes to edit the band-tailed pigeon, the passenger pigeon’s closest living relative, with as many of the extinct bird’s defining traits as possible. Eventually, he says, he’ll have a hybrid creature that looks and acts like a passenger pigeon (albeit with no parental training) but still contains band-tailed pigeon DNA. These new-old birds will need a name, which their human creator has already chosen: Patagioenas neoectopistes, or “new wandering pigeon of America.”
Excerpts from Amy Dockser Marcus, Meet the Scientists Bringing Extinct Species Back From the Dead, WSJ, the Future of Everything, Oct. 11, 2018
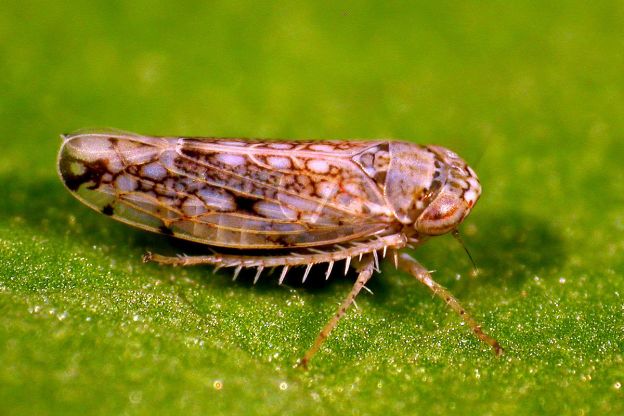

According to Science Magazine, Agricultural genetic technologies typically achieve their agronomic aims by introducing laboratory-generated modifications into target species’ chromosomes. However, the speed and flexibility of this approach are limited, because modified chromosomes must be vertically inherited from one generation to the next. In an effort to remove this limitation, an ongoing research program funded by the U.S. Defense Advanced Research Projects Agency (DARPA) aims to disperse infectious genetically modified viruses that have been engineered to edit crop chromosomes directly in fields [through insects]. This is genetic engineering through horizontal transfer, as opposed to vertical inheritance. The regulatory, biological, economic, and societal implications of dispersing such horizontal environmental genetic alteration agents (HEGAAs)[eg leafhoppers, whiteflies and aphids) into ecosystems are profound. Further, this program stipulates that the means of delivery of these viral HEGAAs into the environment should be insect-based dispersion (Insect Allies Program). In the context of the stated aims of the DARPA program, it is our opinion that the knowledge to be gained from this program appears very limited in its capacity to enhance U.S. agriculture or respond to national emergencies (in either the short or long term). Furthermore, there has been an absence of adequate discussion regarding the major practical and regulatory impediments toward realizing the projected agricultural benefits. As a result, the program may be widely perceived as an effort to develop biological agents for hostile purposes and their means of delivery, which—if true—would constitute a breach of the Biological Weapons Convention (BWC).

It’s an eye-catching statistic: A single company, the multinational chemical giant BASF, owns nearly half of the patents issued on 13,000 DNA sequences from marine organisms. That number is now helping fuel high-stakes global negotiations on a contentious question: how to fairly regulate the growing exploitation of genes collected in the open ocean, beyond any nation’s jurisdiction.
The negotiations that took place at the UN in September 2018 aim, inter alia, to replace today’s free-for-all scramble for marine genetic resources with a more orderly and perhaps more just regime. Many developed nations and industry groups are adamant that any new rules should not complicate efforts to discover and patent marine genes that may help create better chemicals, cosmetics, and crops. But many developing nations want rules that will ensure they, too, share in any benefits. Scientists are also watching. A regulatory regime that is too burdensome could have “a negative impact” on scientists engaged in “noncommercial ocean research,” warns Robert Blasiak, a marine policy specialist at the Stockholm Resilience Centre. It is not the first time nations have wrangled over how to share genetic resources. Under another U.N. pact, the 2010 Nagoya Protocol, 105 countries have agreed to rules to prevent so-called biopiracy: the removal of biological resources—such as plant or animal DNA—from a nation’s habitats without proper permission or compensation.
Those rules don’t apply in international waters, which begin 200 nautical miles from shore and are attracting growing interest from researchers and companies searching for valuable genes. The first patent on DNA from a marine organism was granted in 1988 for a sequence from the European eel, which spends part of its life in freshwater. Since then, more than 300 companies, universities, and others have laid claim to sequences from 862 marine species, a team led by Blasiak reported in June in Science Advances. Extremophiles have been especially prized. Genes from worms found in deep-sea hydrothermal vents, for example, encode polymers used in cosmetics. And BASF has patented other worm DNA that the company believes could help improve crop yields. The conglomerate, based in Ludwigshafen, Germany, says it found most of its 5700 sequences in public databases…
It may take years for nations to agree on a marine biodiversity treaty; [A]n “ideological divide” between developing and developed countries has, so far, “led to stalemate” on how to handle marine genetic resources, says Harriet Harden-Davies, a policy expert at the University of Wollongong in Australia.
Most developing nations want to expand the “common heritage” philosophy embedded in the 1982 United Nations Convention on the Law of the Sea, which declares that resources found on or under the seabed, such as minerals, are the “common heritage of mankind.” Applying that principle to genetic resources would promote “solidarity in the preservation and conservation of a good we all share,” South Africa’s negotiating team said in a recent statement. Under such an approach, those who profit from marine genes could, for example, pay into a global fund that would be used to compensate other nations for the use of shared resources, possibly supporting scientific training or conservation.
But developed nations including the United States, Russia, and Japan oppose extending the “common heritage” language, fearing burdensome and unworkable regulations. They argue access to high seas genes should be guaranteed to all nations under the principle of the “freedom of the high seas,” also enshrined in the Law of the Sea. That approach essentially amounts to finders keepers, although countries traditionally have balanced unfettered access with other principles, such as the value of conservation, in developing rules for shipping, fishing, and research in international waters.
The European Union and other parties want to sidestep the debate and seek a middle ground. One influential proposal would allow nations to prospect for high seas genes, but require that they publish the sequences they uncover. Companies could also choose to keep sequences private temporarily, in order to be able to patent them, if they contribute to an international fund that would support marine research by poorer nations. “Researchers all around the world should be put all on a level playing field,” says Arianna Broggiato, a Brussels-based legal adviser for the consultancy eCoast, who co-authored a paper on the concept this year in The International Journal of Marine and Coastal Law.
Exceprts from Eli Kintisch U.N. tackles gene prospecting on the high seas, Science, Sept. 7, 2018

 The world’s vast oceans and seas offer seemingly endless spaces in which adversaries of the United States can maneuver undetected. The U.S. military deploys networks of manned and unmanned platforms and sensors to monitor adversary activity, but the scale of the task is daunting and hardware alone cannot meet every need in the dynamic marine environment. Sea life, however, offers a potential new advantage. Marine organisms are highly attuned to their surroundings—their survival depends on it—and a new program out of DARPA’s Biological Technologies Office aims to tap into [marine animals] natural sensing capabilities to detect and signal when activities of interest occur in strategic waters such as straits and littoral regions.
The world’s vast oceans and seas offer seemingly endless spaces in which adversaries of the United States can maneuver undetected. The U.S. military deploys networks of manned and unmanned platforms and sensors to monitor adversary activity, but the scale of the task is daunting and hardware alone cannot meet every need in the dynamic marine environment. Sea life, however, offers a potential new advantage. Marine organisms are highly attuned to their surroundings—their survival depends on it—and a new program out of DARPA’s Biological Technologies Office aims to tap into [marine animals] natural sensing capabilities to detect and signal when activities of interest occur in strategic waters such as straits and littoral regions.
The Persistent Aquatic Living Sensors (PALS) program, led by program manager Lori Adornato, will study natural and modified organisms to determine which ones could best support sensor systems that detect the movement of manned and unmanned underwater vehicles. PALS will investigate marine organisms’ responses to the presence of such vehicles, and characterize the resulting signals or behaviors so they can be captured, interpreted, and relayed by a network of hardware devices.
Beyond sheer ubiquity, sensor systems built around living organisms would offer a number of advantages over hardware alone. Sea life adapts and responds to its environment, and it self-replicates and self-sustains. Evolution has given marine organisms the ability to sense stimuli across domains—tactile, electrical, acoustic, magnetic, chemical, and optical. Even extreme low light is not an obstacle to organisms that have evolved to hunt and evade in the dark.
However, evaluating the sensing capabilities of sea life is only one of the challenges for PALS researchers. Performer teams supporting DARPA will also have to develop hardware, software, and algorithms to translate organism behavior into actionable information and then communicate it to end users…. The complete sensing systems must also discriminate between target vehicles and other sources of stimuli, such as debris and other marine organisms, to limit the number of false positives.
Adornato is aiming to demonstrate the approach and its advantages in realistic environments to convey military utility. “Our ideal scenario for PALS is to leverage a wide range of native marine organisms, with no need to train, house, or modify them in any way, which would open up this type of sensing to many locations,” Adornato said.
Excerpt from PALS Turns to Marine Organisms to Help Monitor Strategic Waters: Highly adapted sea life could help U.S. military detect adversary activity over large areas, Feb. 2, 2018
Consider the benefits to be gained from a chimney that heals after damage, a roof that breathes to control airflow, surfaces that don’t flake or fade, and a driveway that eats oil to clean up after spills.–From the DARPA website
The structural materials that are currently used to construct homes, buildings, and infrastructure are expensive to produce and transport, wear out due to age and damage, and have limited ability to respond to changes in their immediate surroundings. Living biological materials—bone, skin, bark, and coral, for example—have attributes that provide advantages over the non-living materials people build with, in that they can be grown where needed, self-repair when damaged, and respond to changes in their surroundings. The inclusion of living materials in human-built environments could offer significant benefits; however, today scientists and engineers are unable to easily control the size and shape of living materials in ways that would make them useful for construction.
DARPA is launching the Engineered Living Materials (ELM) program with a goal of creating a new class of materials that combines the structural properties of traditional building materials with attributes of living systems. Living materials represent a new opportunity to leverage engineered biology to solve existing problems associated with the construction and maintenance of built environments, and to create new capabilities to craft smart infrastructure that dynamically responds to its surroundings.
“The vision of the ELM program is to grow materials on demand where they are needed,” said ELM program manager Justin Gallivan. “Imagine that instead of shipping finished materials, we can ship precursors and rapidly grow them on site using local resources. And, since the materials will be alive, they will be able to respond to changes in their environment and heal themselves in response to damage.”…
Scientists are making progress with three-dimensional printing of living tissues and organs, using scaffolding materials that sustain the long-term viability of the living cells. These cells are derived from existing natural tissues, however, and are not engineered to perform synthetic functions. And current cell-printing methods are too expensive to produce building materials at necessary scales.
ELM looks to merge the best features of these existing technologies and build on them to create hybrid materials composed of non-living scaffolds that give structure to and support the long-term viability of engineered living cells. DARPA intends to develop platform technologies that are scalable and generalizable to facilitate a quick transition from laboratory to commercial applications.
The long-term objective of the ELM program is to develop an ability to engineer structural properties directly into the genomes of biological systems so that neither scaffolds nor external development cues are needed for an organism to realize the desired shape and properties. ….
Work on ELM will be fundamental research carried out in controlled laboratory settings. DARPA does not anticipate environmental release during the program.
Excerpts from Living Structural Materials Could Open New Horizons for Engineers and Architects, DARPA seeks to develop design tools and methods for creating programmable, self-healing, living building materials, OUTREACH@DARPA.MIL, Aug. 5, 2016
See also FBO.org
From the DARPA website:
The development of increasingly sophisticated techniques and tools to sequence, synthesize and manipulate genetic material has led to the rapidly maturing discipline of synthetic biology. …[But] The costs of maintaining required environmental controls and detecting and compensating for genetic alterations are substantial and severely limit the widespread application of synthetic biology to U.S. national security missions.
To help address these challenges, DARPA has created the Biological Robustness in Complex Settings (BRICS) BRICS seeks to develop the fundamental understanding and component technologies needed to increase the biological robustness and stability of engineered organisms while maintaining or enhancing the safe application of those organisms in complex biological environments. The goal is to create the technical foundation for future engineered biological systems to achieve greater biomedical, industrial and strategic potential.
“By making these systems more robust, stable and safe, BRICS seeks to harness the full range of capabilities at the intersection of engineering and biology,” said Justin Gallivan, DARPA program manager. “These capabilities could include efficient on-demand bio-production of novel drugs, fuels, sensors and coatings; or engineered microbes able to optimize human health by treating or preventing disease.”
Excerpt from BUILDING THE FOUNDATION FOR FUTURE SYNTHETIC BIOLOGY APPLICATIONS WITH BRICS, July 29, 2014